Note
Click here to download the full example code
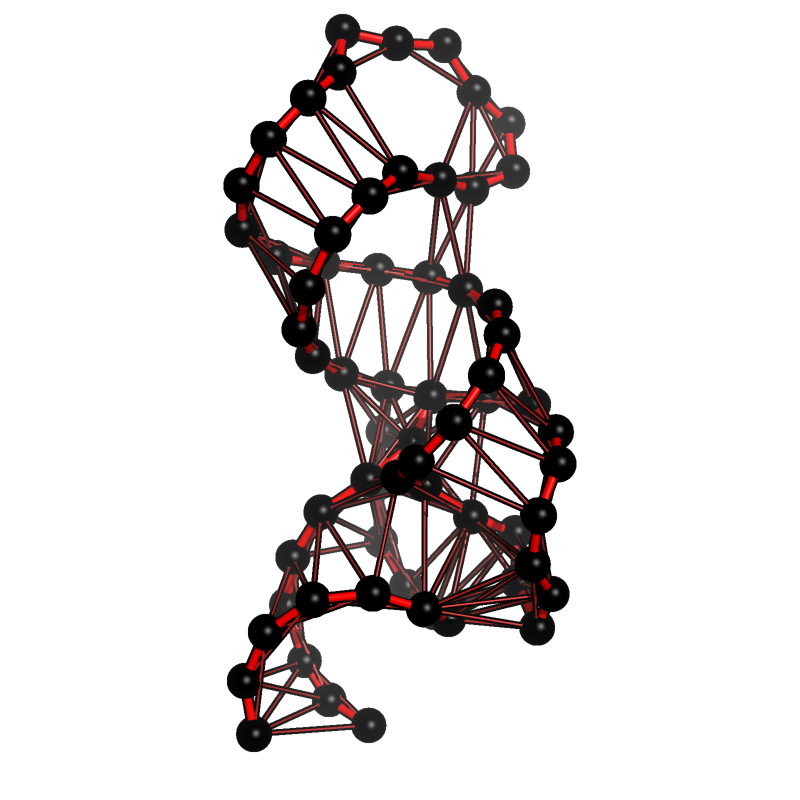
Contacts between nucleotides in a tetracycline aptamer¶
This example reproduces a figure from the publication “StreAM-Tg: algorithms for analyzing coarse grained RNA dynamics based on Markov models of connectivity-graphs” 1.
The figure displays a coarse grained model of a tetracycline aptamer and highlights interacting nucleotides based on a cutoff distance.
- 1
S Jager, B Schiller, P Babel, M Blumenroth, T Strufe and K Hamacher, “StreAM-Tg: algorithms for analyzing coarse grained RNA dynamics based on Markov models of connectivity-graphs.” Algorithms Mol Biol 12 (2017).
# Code source: Patrick Kunzmann
# License: CC0
import numpy as np
import biotite.structure as struc
import biotite.structure.io.mmtf as mmtf
import biotite.database.rcsb as rcsb
import ammolite
PNG_SIZE = (800, 800)
mmtf_file = mmtf.MMTFFile.read(rcsb.fetch("3EGZ", "mmtf"))
structure = mmtf.get_structure(mmtf_file, model=1)
aptamer = structure[struc.filter_nucleotides(structure)]
# Coarse graining: Represent each nucleotide using its C3' atom
aptamer = aptamer[aptamer.atom_name == "C3'"]
# Connect consecutive nucleotides
indices = np.arange(aptamer.array_length())
aptamer.bonds = struc.BondList(
aptamer.array_length(),
np.stack((indices[:-1], indices[1:]), axis=-1)
)
pymol_obj = ammolite.PyMOLObject.from_structure(aptamer)
pymol_obj.show("sticks")
pymol_obj.show("spheres")
pymol_obj.color("black")
ammolite.cmd.set("stick_color", "red")
ammolite.cmd.set("stick_radius", 0.5)
ammolite.cmd.set("sphere_scale", 1.0)
ammolite.cmd.set("sphere_quality", 4)
# Adjust camera
pymol_obj.orient()
pymol_obj.zoom(buffer=10)
ammolite.cmd.rotate("z", 90)
ammolite.show(PNG_SIZE)

CUTOFF = 13
# Find contacts within cutoff distance
adjacency_matrix = struc.CellList(aptamer, CUTOFF) \
.create_adjacency_matrix(CUTOFF)
for i, j in zip(*np.where(adjacency_matrix)):
pymol_obj.distance("", i, j, show_label=False, gap=0)
ammolite.cmd.set("dash_color", "firebrick")
# Add black outlines
ammolite.cmd.bg_color("white")
ammolite.cmd.set("ray_trace_mode", 1)
ammolite.cmd.set("ray_trace_disco_factor", 0.5)
ammolite.show(PNG_SIZE)
# sphinx_gallery_thumbnail_number = 2